Attachments:Hi Pymolers,I need to perform a batch operation but I need a help with scriptmodification.A have a set (100) of pdb files that I would like to mutate all residues tolysine. I modified mutate02.py script in pymol to perform this task, but Iwould need two extra 'features':1) the script would need to save the file with a different name. Forexample, load test.pdb, mutate all residues to lysine and then save themutated version to testK.pdb2) the script would need to perform this task for all the pdb files in adirectory.Right now, I am not concerned about the possible rotamers of lysine, anylysine rotamer would work.Could anyone give a hint on how to do it?Thanks!Fabio.Thread view. Attachments:Hi Pymolers,I need to perform a batch operation but I need a help with scriptmodification.A have a set (100) of pdb files that I would like to mutate all residues tolysine. I modified mutate02.py script in pymol to perform this task, but Iwould need two extra 'features':1) the script would need to save the file with a different name.
Forexample, load test.pdb, mutate all residues to lysine and then save themutated version to testK.pdb2) the script would need to perform this task for all the pdb files in adirectory.Right now, I am not concerned about the possible rotamers of lysine, anylysine rotamer would work.Could anyone give a hint on how to do it?Thanks!Fabio.
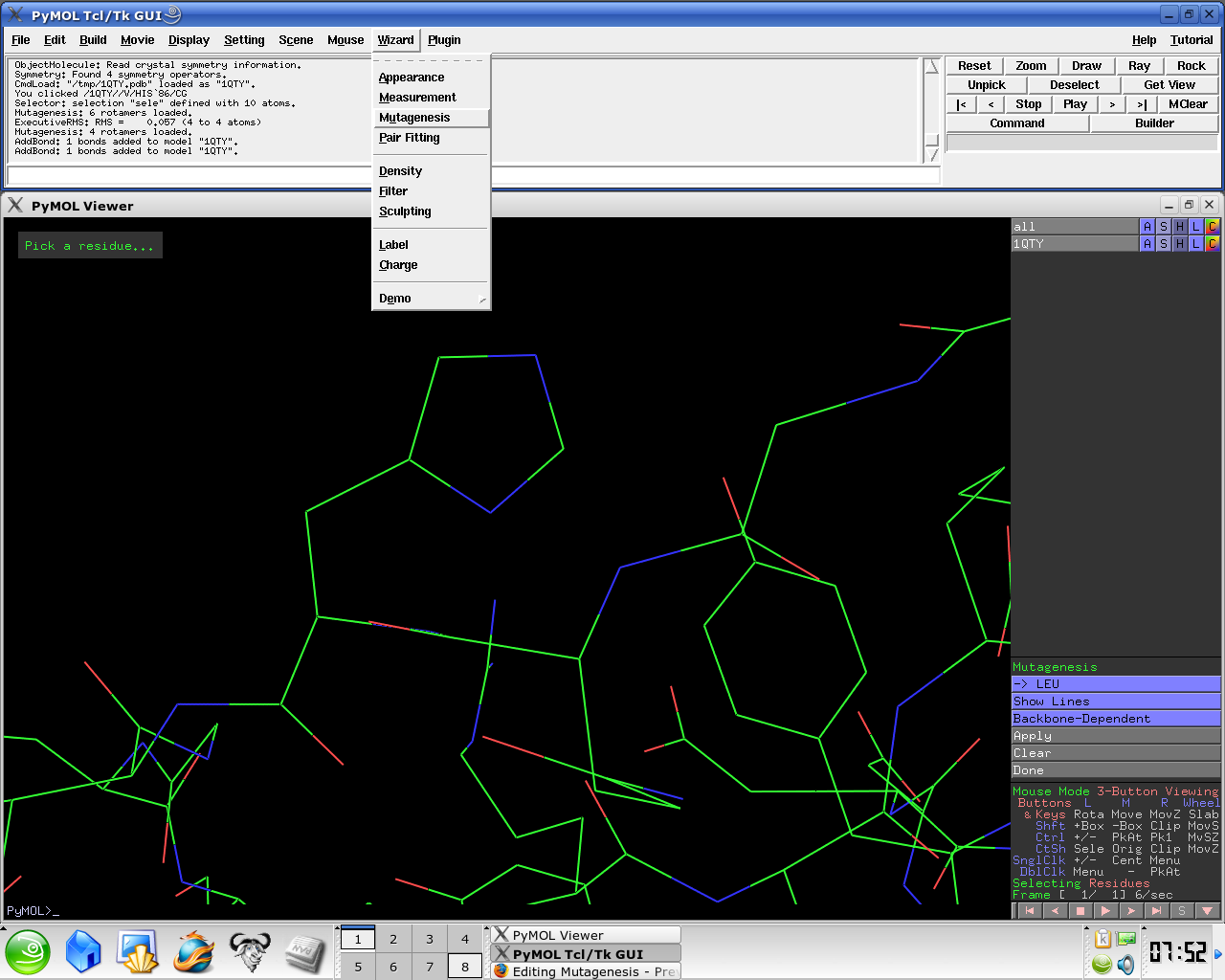
Mutating Amino-Acids. To initiate a mutation, simply click on the mutation tool located at the top of the display window. You will be prompted to select a group, which is readily made by clicking on any atom of the desired amino acid. Then a pop-up menu will let you select the new amino acid type. Question: C Api To Mutate Pdb Residues. Either way, once you have your PDB data parsed into memory, you might write a mutate function that iterates through and operates on instances of the desired PDB record type in your dataset, to change them per your specification. Or insert and delete functions that add or remove an instance.
